Habitat Suitability Modelling
Conservation planning depends on understanding where species can survive and thrive.
Traditional approaches to habitat suitability modelling require extensive human input throughout the process. Collecting and processing environmental layers and species observation data and selecting variables before any modelling can begin. It can be an iterative and slow process. This project developed a comprehensive platform that automates the entire workflow, freeing up the time of subject matter experts to do what they do best.
The work began as my M.Sc. dissertation at Imperial College London, investigating whether transformer models could be applied to habitat prediction. The results earned a distinction and demonstrated that deep learning approaches could match traditional statistical methods. Development continued at AI For Nature Lab, exploring alternative architectures and incorporating species range maps, then expanded significantly at WildMon.
The current platform integrates multiple modelling approaches including TabNet, XGBoost, and random forests. It connects directly to Google Earth Engine for automated satellite and environmental data collection globally, pulls species occurrence records from the Global Biodiversity Information Facility. It allows users to incorporate local files from eDNA, Bioacoustics and other sources and includes comprehensive data processing, validation, and visualisation tools. A user defines their study area as a polygon anywhere on Earth, and the system handles everything else.
The platform’s most significant application was a habitat suitability study for chimpanzees in Mali, delivered on behalf of WildMon for the Jane Goodall Institute, on which I served as lead author.
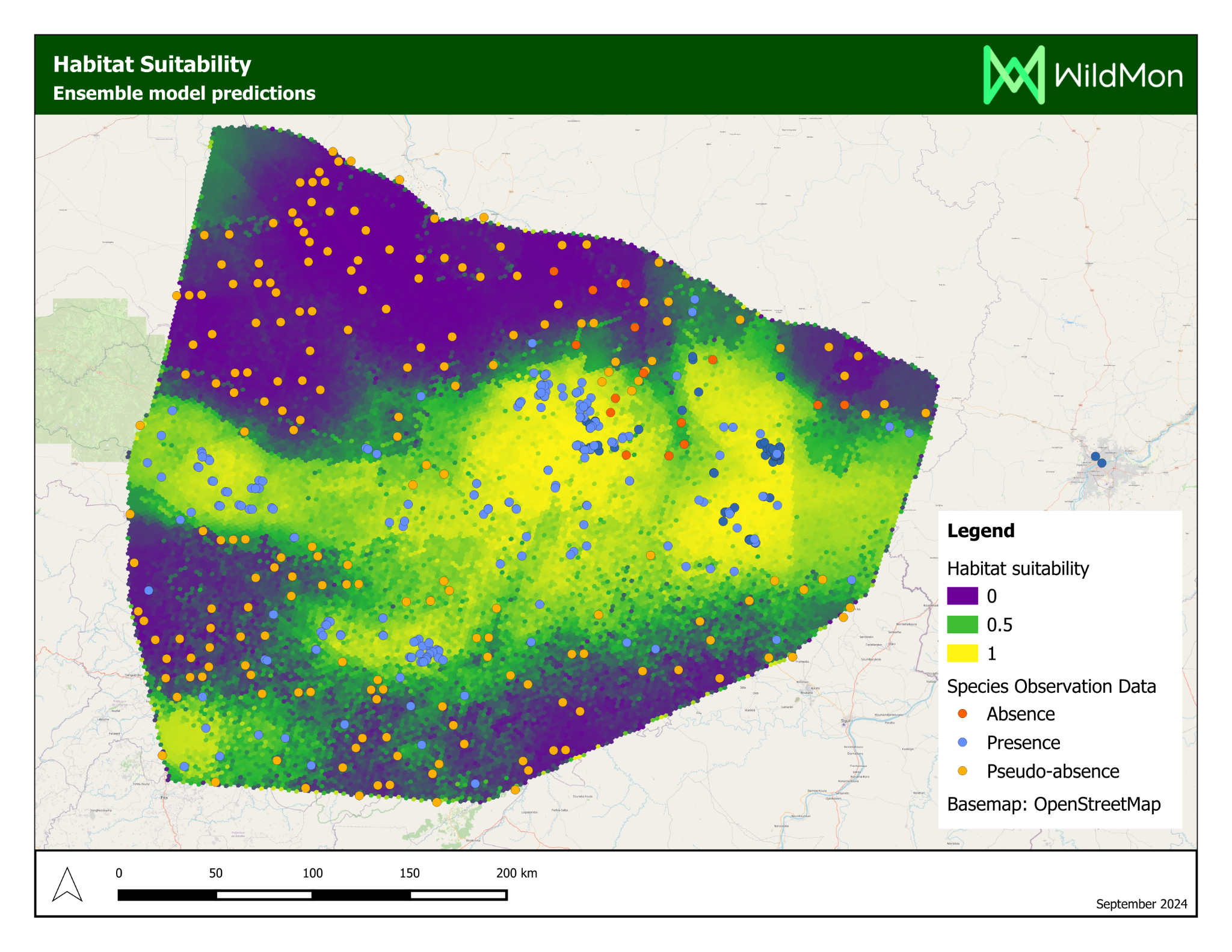
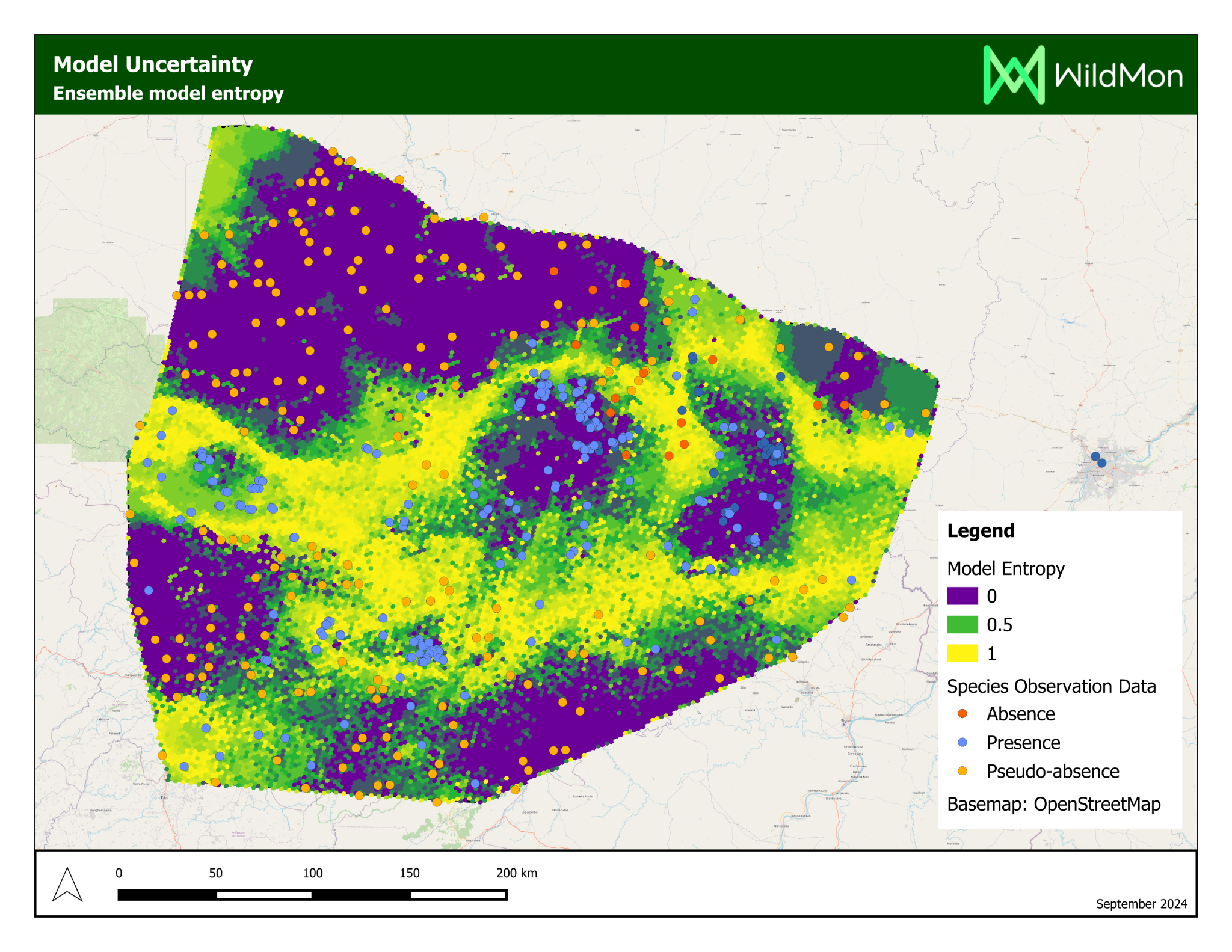
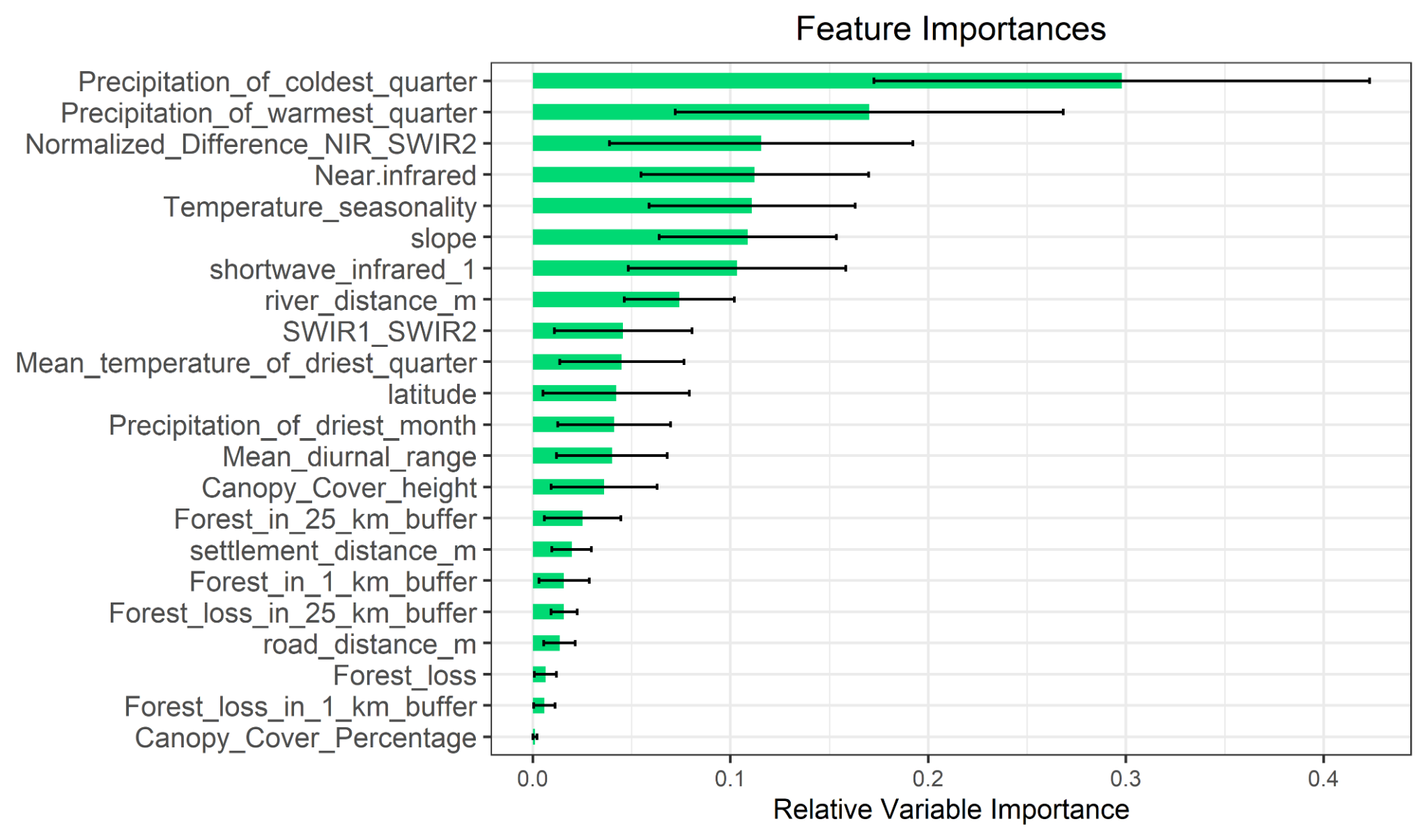

Project Partners



